Antibody molecules can often, but not always, recover from extensive denaturation by urea, guanidine and extremes of pH (see also Chapter 11) Polyclonal antisera generally require quite harsh conditions (6–8 M urea, 5–6 M guanidinehydrochloride, pH < 3 or > 10) before the antigen–antibody bond is disrupted (1930) Denaturation of proteins by urea and related substances Nature 126 328 – 330, 3 – 384 OpenUrl CrossRef ↵ Greene RF, Jr, Pace CN (1974) Urea and guanidine hydrochloride denaturation of ribonuclease, lysozyme,Protein denaturation with guanidine hydrochloride or urea provides a different estimate of stability depending on the contributions of electrostatic interactions O D Monera, C M Kay, and R S Hodges Author informationCopyright and License informationDisclaimer

Preparation And Extraction Of Insoluble Inclusion Body Proteins From Escherichia Coli Abstract Europe Pmc
How does guanidine hydrochloride denature proteins
How does guanidine hydrochloride denature proteins-The inactivation of papain brought about by low concentration of urea or guanidine hydrochloride was found to be a noncompetitive type and that of cyanate ion to be a and hydrogen bonds in the protein mole cule1†`3) These studies, in connection with changes of denaturation of these enzyme mole cules due to denaturing reagent (FigsGuanidine HCl is a freely soluble chaotropic agent routinely used for protein denaturation and folding In order to study their molecular properties and amino acid composition, proteins must be denatured from their folded state The exact mechanism of protein unfolding by guanidine HCl is unclear However it is speculated that the denaturant migrates into the interior of the protein
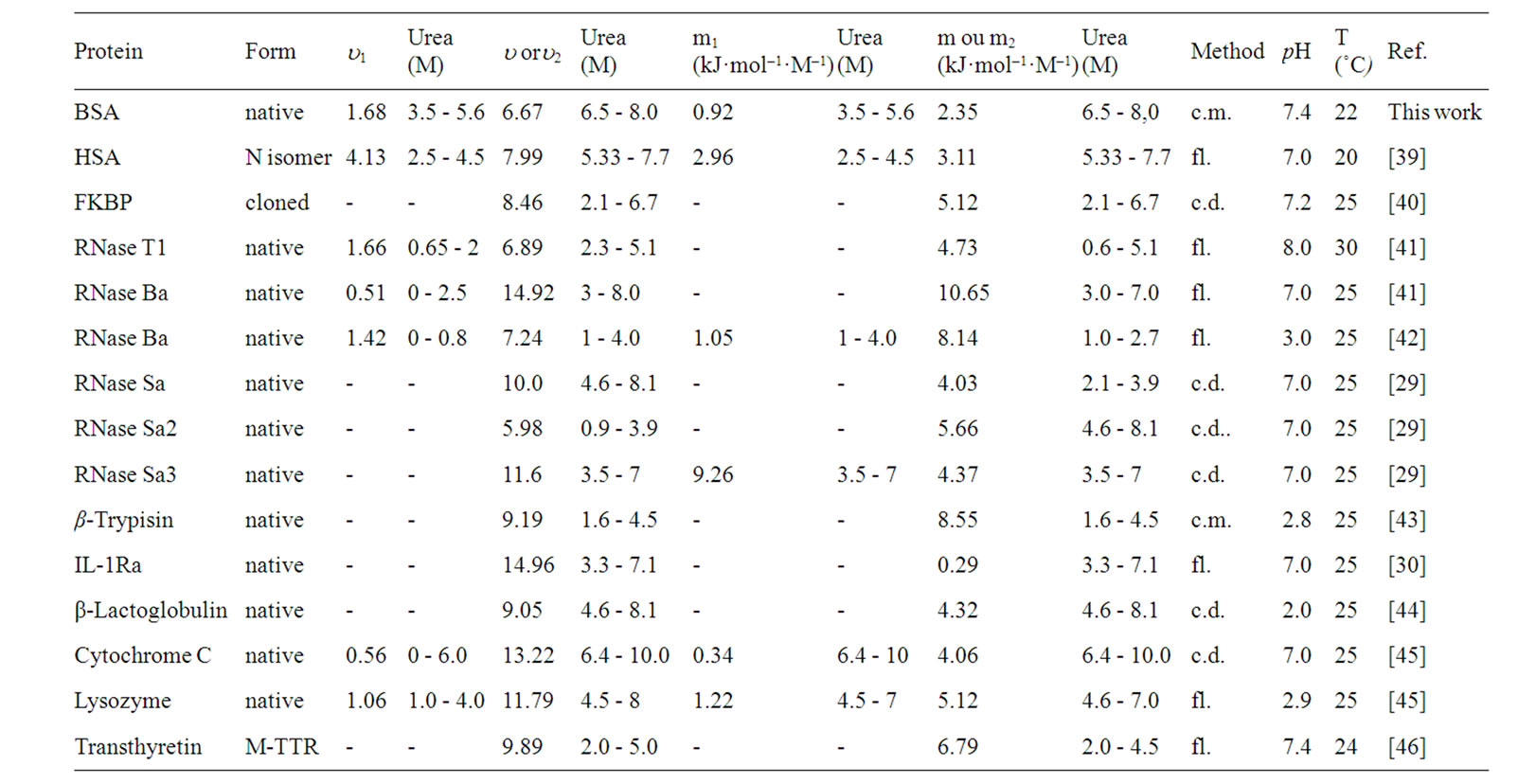



Pressure And Urea Induced Denaturation Of Bovine Serum Albumin Considerations About Protein Heterogeneity
Guanidine hydrochloride (GdnHCl) give the same estimates of the stability of a particular protein Moreover, estimates of protein stability from GdnHCl and urea denaturation data might differ11 rows Protein denaturation using guanidineHCl and urea (and often temperature) are monitored Changing pH denatures proteins because it changes the charges on many of the side chains This disrupts electrostatic attractions and hydrogen bonds ii Certain reagents such as urea and guanidine hydrochloride denature proteins by forming hydrogen bonds to the protein groups that are stronger than the hydrogen bonds formed between the groups
In general, as commonly used reagents in protein denaturation, guanidine hydrochloride has relatively strong dissolving capacity and denatured ability and does not cause the covalent modification of the recombinant protein, but has the disadvantages of high cost, easy precipitation under acidic conditions, and interference with protein ion exchange chromatography;Guanidine HydrochlorideGuanidine chloridepromoted protein denaturation can be used in protein purification and analysis as well as isolation of proteinbound ligands Storage Room temperature Keywords Aminoformamidine hydrochloride, GuHCl, Guanidinium chloride, Guanidium chloride Grade Biotechnology grade All components are highly pure (minimum 99%) An undergraduate laboratory experiment to measure the free energy of denaturation of the enzyme lysozyme is described The fluorescence of the protein in solutions of varying concentrations of urea and guanidine hydrochloride is measured The data are analyzed graphically assuming a twostate model for the denaturation
Protein unfolding induced by chemical denaturants such as urea and guanidine hydrochloride (GdnHCl) is a common approach to study protein folding in vitro Meanwhile, it has been shown that low concentrations of GdnHCl can cause protein stabilization by eliminating the strains in protein caused by the electrostatic interactions of charged groups on its surface 2 , 3Of proteins with high content of βstructure and, especially, proteins with βbarrel topology is significantly less understood with respect to folding of αhelical and α/β proteins This work is focused on investigation of denaturation processes induced by guanidine hydrochloride (GdnHCl) of proteins posThe thermal denaturation is demonstrated to be twostate and reversible upon the addition of low concentrations of added guanidine hydrochloride (GuHCl) DG values from the DSC studies are in excellent agreement with values from isothermal GuHCl denaturation monitored by fluorescence and circular dichroism (CD) spectroscopy
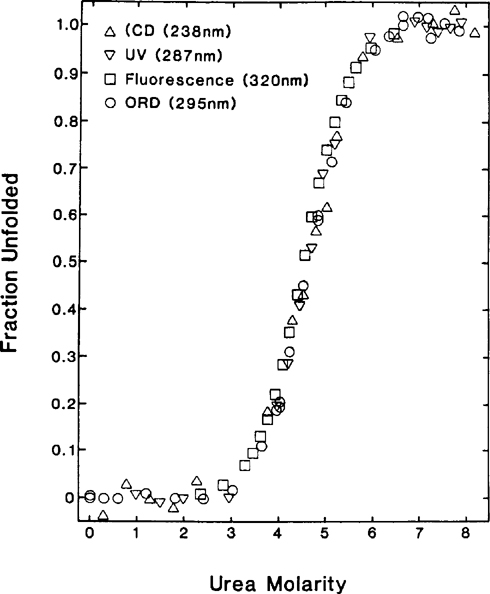



Urea And Guanidine Hydrochloride Denaturation Curves Springerlink



2
KeywordsChymotrypsin, denaturation, osmolytes, protease inhibitor, serine proteases, turkey ovomucoid third domain AbstractIn this communication we describe an approach in which guanidine hydrochlorideinduced dissociation of a protein inhibitorserine protease complex is used to explore the molecular basis of protein denaturation TheEfficient protein denaturation with consequent elimination of nucleolytic damage to the RNA requires rapid and complete homogenization For instance, homogenization of a rat pancreas in guanidinium thiocyanate solution by means of a motorized pestle results in extensively degraded RNA, which is avoided by the use of an apparatus such as aDenaturation is the physical or chemical effect on proteins which result in its structure and property change The change and deterioration of secondary and tertiary structures for the protein show its denaturation Some reagents, such as guanidine hydrochloride and urea, will make protein denaturize The reagents are known as denaturants




Adapting The Chemical Unfolding Assay For High Throughput Protein Screening Using Experimental And Spectroscopic Corrections Sciencedirect




Protein Denaturation And Misfolding
Web http//wwweurekaselectcom/1057/articleTitle Investigation of the Mechanism of Protein Denaturation by Guanidine HydrochlorideInduced Dissociation The activity and conformational change of human placental cystatin (HPC), a low molecular weight thiol proteinase inhibitor (12,500) has been investigated in presence of guanidine hydrochloride (GdnHCl) and urea The denaturation of HPC was followed by activity measurements, fluorescence spectroscopy and Circular Dichroism (CD) studiesBiochem/physiol Actions Guanidine hydrochloride (GuHCl) is a small hydroscopic molecule It plays a role in inhibiting heat shock protein 104 (Hsp104) adenosine triphosphatase (ATPase) activity in vivo It is a potent denaturant and inactivator of several enzymes and proteins GuHCl can inactivate aminoacylase and papain
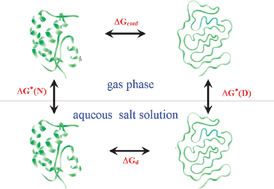



Contrasting The Denaturing Effect Of Guanidinium Chloride With The Stabilizing Effect Of Guanidinium Sulfate Physical Chemistry Chemical Physics Rsc Publishing




Pressure And Urea Induced Denaturation Of Bovine Serum Albumin Considerations About Protein Heterogeneity
Guanidine hydrochloride (GdnHCl) denaturation using circular dichroism, fluorescence and UVdifference spectroscopy Buffers used in these experiments were 005 M sodium phosphate buffer, pH 75, 015 M TrisHCl buffer, pH 75, 015 M HEPES buffer, pH 75 and 015 M MOPS buffer, pH 75 Differential effects ofIn general, proteins require denaturation and disulfide bond cleavage for enzymatic digestion to reach completion If digestion of a native protein is desired, begin this protocol at Step 3 Dissolve the target protein in 6M guanidine HCl (or 8M urea), 50mM TrisHCl (pH 8), 2–5mM DTT Heat at 37°C for 45–60 minutesGuanidine hydrochloride ≥99,7 %, pa, protein Grade Chaotropic agent that potently destroys the threedimensional structure of proteins, leading to denaturation Guanidin hydrochloride is used for protein denaturation as well as for releasing proteins from fusion bodies (Phage plaque membrane filters), and for renaturation/refolding of proteins



Denaturation Of Proteins In Guanidine Hcl Fluorescence Of 0 3 M Download Scientific Diagram




Guanidine Hydrochloride Bioultra For Molecular Biology 99 5 At 50 01 1
Consequently, some proteins will be irreversibly altered upon interaction with guanidine solutions and may lose their binding function Before any largescale use of guanidine, it is best to test a small sample and determine whether the denaturing effects will adversely affect the intended use of the protein Related Products 8M GuanidineHClAbbreviations PDI, protein disulfide isomerase;Guanidine HCl was purified according to the method of Reithel and Sakura (14) by recrystallization from methanol The molar extinction at 233 urn was less than 002 11l cml The samples with guanidine HCl were prepared in 3ml volu metric flasks, the solutions were adjusted to




Quantitating Denaturation By Formic Acid Imperfect Repeats Are Essential To The Stability Of The Functional Amyloid Protein Fapc Biorxiv




The Stability Of A Three State Unfolding Protein Intechopen
Protein analysis methods including mass spectrometry, electrophoresis, and enzymatic digests Guanidine hydrochloride has typically been used for the isolation of RNA, to denature globular proteins, and for protein refolding studies It can also be used to facilitate the generation of tryptic peptides for analysis of complex protein samples Guanidine hydrochloride(CAS) is commonly used as an intermediate in pharmaceuticals, pesticides, dyes and other organic synthetics, and is often used in denaturing proteins, as well as in refolding proteins after denaturation Guanidine hydrochloride supplier will explain the role of guanidine hydrochloride in protein denaturation and refoldingProtocol for Guanidinium Denaturation Studies I Make Stock Solutions A Make ~ 75 M Gua·HCl titration 1 Dissolve 358 g Gua·HCl in ~30 mL water 2 Volume will expand 3 Add small amounts of water until solid is entirely dissolved 4 Bring volume up to 500 mL 5 Measure concentration of solution by refractive index measurements (Pace CN, Meth
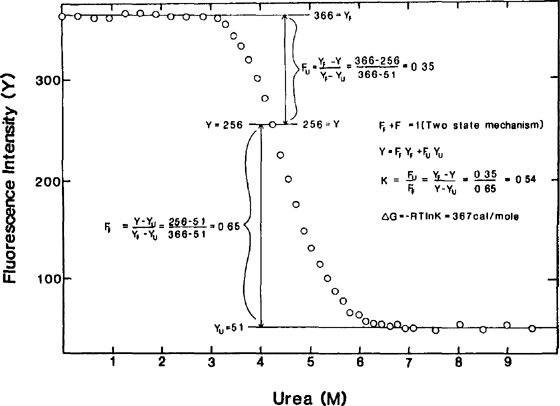



Urea And Guanidine Hydrochloride Denaturation Curves Springerlink
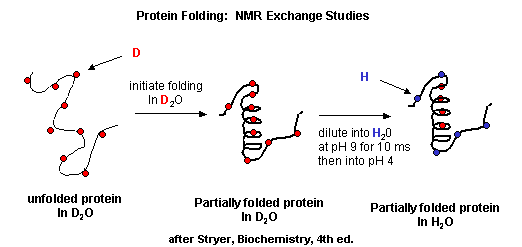



D2 Protein Folding In Vitro Biology Libretexts
Consequently, some proteins will be irreversibly altered upon interaction with guanidine solutions and may lose their binding function Before any largescale use of guanidine, it is best to test a small sample and determine whether the denaturing effects will adversely affect the intended use of the protein Related Products GuanidineHClThe denaturation of the dimeric enzyme glucose6phosphate dehydrogenasc from Leuconostoc mesenteroides by guanidine hydrochloride has been studied using enzymatic activity, intrinsic fluorescence, circular dichroism, and light scattering measure mentsProtein denaturation with guanidine hydrochloride or urea provides a different estimate of stability depending on the contributions of electrostatic interactions The objective of this study was to address the question of whether or not urea and guanidine hydrochloride (GdnHCl) give the same estimates of the stability of a particular protein We previously suspected that the




Kinetic Evidence For A Two Stage Mechanism Of Protein Denaturation By Guanidinium Chloride Pnas




Pdf Effect Of Chaotropic Reagents On Bovine Serum Albumin Bsa A Fluorescence Study Semantic Scholar
The denaturation and renaturation process of pyruvate decarboxylase (PDC) from Zymomonas mobilis (ATCC ) has been investigated using guanidine hydrochloride and urea as denaturing agents The quarternary structure of the homotetramer is strongly stabilized by the cofactors Mg2 and thiamine diphosphate (TDP)Guanidine hydrochloride, denaturant, chaotrope A selection of strong chemicals to aid in the denaturation of proteins Denaturants, or chaotropes, disrupt water interactions resulting in the solubilization of hydrophobic proteins and peptides Hi everybody, I am supposed to do a denaturation of my protein in 6M guanidinhydrochloride followed by several wash steps in decreasing concentrations of guanidine hcl the protein is in this case already fixated on nitrocellulose and is going to be subject to immunodetection afterwards



1




Denaturation Of The Wild Type And Mutant Bgltm Proteins With Guanidine Hydrochloride
Protein Denaturation Urea vs Guanidine Hydrochloride (GdnHCl) In equilibrium denaturation experiments, the concentration of urea at midpoint of denaturation is always higher than that of Denaturation of EGFP or sfGFP induced by guanidine hydrochloride (GdnHCl) achieves quasiequilibrium after several days of protein incubation in the presence of this denaturing agent Figure 3 shows the kinetics of the denaturation of sfGFP induced by GdnHCl and guanidine thiocyanate (GTC), as recorded by green chromophore fluorescence In theProtein denaturation with guanidine hydrochloride or urea provides a different estimate of stability depending on the contributions of electrostatic interactions Oscar D Monera Department of Biochemistry and the Protein Engineering Network of Centres of Excellence, University of Alberta, Edmonton, Alberta T6G 2H7, Canada



160



2
Guanidine denaturation analysis showed that RAP has two phases in its denaturation, an early denaturation transition at 06 m guanidine HCl, and a broad second transition between 10 and 30 m guanidine HCl Analysis of the denaturation of the individual repeats showed that, despite the similarity in sequence and protease sensitivity between Guanidine hydrochloride induced denaturationrenaturation of δ2crystallin The fluorescence transition of δ2crystallin at various concentrations of GdnHCl is shown in Fig 3 The concentration of GdnHCl inducing denaturation of half the protein as measured by fluorescence spectroscopy, GdnHCl 05, was determined to be 16 mThe activity and conformational change of human placental cystatin (HPC), a low molecular weight thiol proteinase inhibitor (12,500) has been investigated in presence of guanidine hydrochloride (GdnHCl) and urea The denaturation of HPC was followed by activity measurements, fluorescence spectroscopy and Circular Dichroism (CD) studies
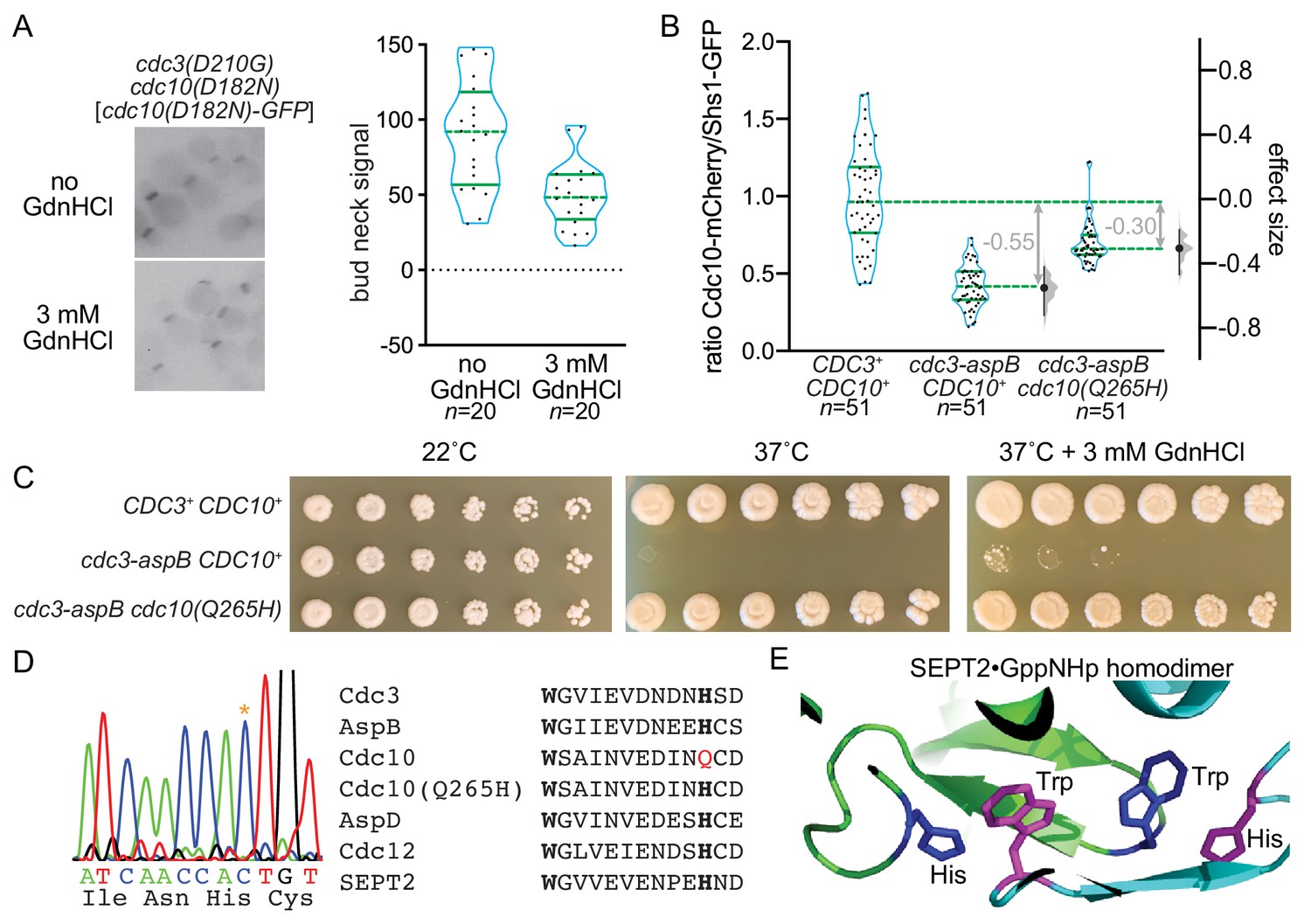



Guanidine Hydrochloride Reactivates An Ancient Septin Hetero Oligomer Assembly Pathway In Budding Yeast Elife




Protein Denaturation With Guanidinium A 2d Ir Study Topic Of Research Paper In Chemical Sciences Download Scholarly Article Pdf And Read For Free On Cyberleninka Open Science Hub
The phenomenon of cold denaturation was investigated for a mAb, mAb1, through thermodynamic and spectroscopic analyses tryptophan fluorescence and circular dichroism (CD) spectra were recorded for the guanidine hydrochloride (GuHCl)induced unfolding of mAb1 at pH 63 at temperatures ranging from 5 to 50°CGdnHCl, guanidine hydrochloride *Towhomreprint requests should be addressed 2107 Thepublication costs ofthis article weredefrayedinpartbypagecharge paymentThisarticle mustthereforebeherebymarked"advertisement" in accordance with 18 USC §1734 solely toThe effect of urea and guanidine hydrochloride (GuHCl) on lysozyme stability has been investigated using activity measurements, microcalorimetry and Raman spectroscopy in the lowfrequency and amide I regions Raman investigations on lysozyme dissolved in H 2 O and D 2 O in the presence of up to 10 M denaturants have revealed direct binding between the protein and




Molecules Free Full Text In Situ Monitored Vortex Fluidic Mediated Protein Refolding Unfolding Using An Aggregation Induced Emission Bioprobe Html




Pdf Urea And Guanidine Hydrochloride Denaturation Of Ribonuclease Lysozyme Zhymotrypsin And Lactoglobulin Semantic Scholar
Summary This chapter contains sections titled Historical Perspective How Urea Denatures Proteins Linear Extrapolation Method ΔG(H2O) mValues Concluding Remarks Experimental Protocols Denaturation of Proteins by Urea and Guanidine Hydrochloride Protein Folding Handbook Wiley Online Library is referred to as the conformational stability of a protein The determinants of native state stability in aqueous solutions are the amino acid sequence of the protein as well as the variable conditions of pH, temperature, and the concentration of salts and ligands (1, 2)Although the native conformation is essential for activity, the conformational stability isGuanidine hydrochloride (GmHCl) as a protein denaturant was first noted by Greenstein in 1938 (11) At present, urea and GmHClr are the most frequently used protein denaturants The main advantage of these denaturants is that the extent of un folding



Protein Denaturation Illuminolist




Guanidinium Chloride Wikipedia
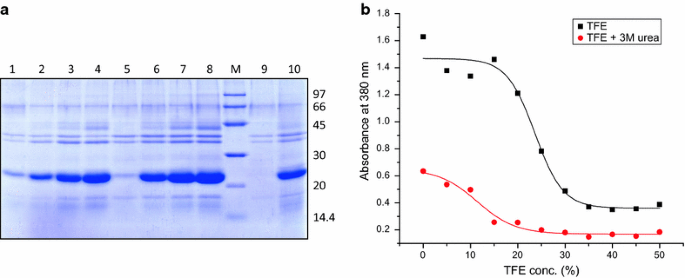



Recovery Of Bioactive Protein From Bacterial Inclusion Bodies Using Trifluoroethanol As Solubilization Agent Microbial Cell Factories Full Text



2
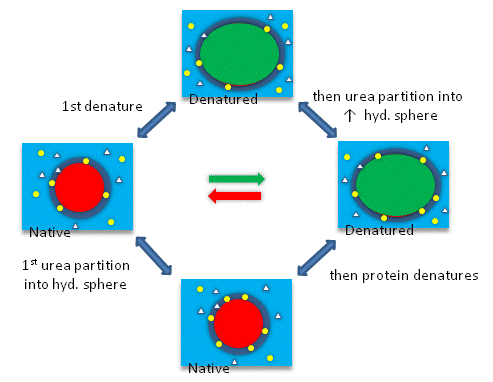



Add Your Page Title
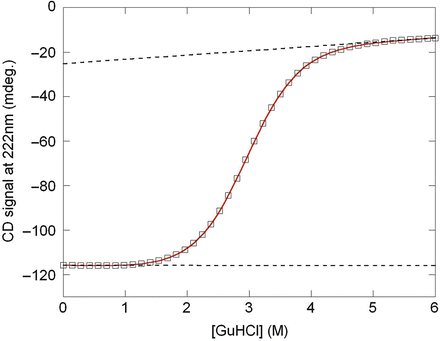



Chemical Denaturation Springerlink



File Yzimgs Com




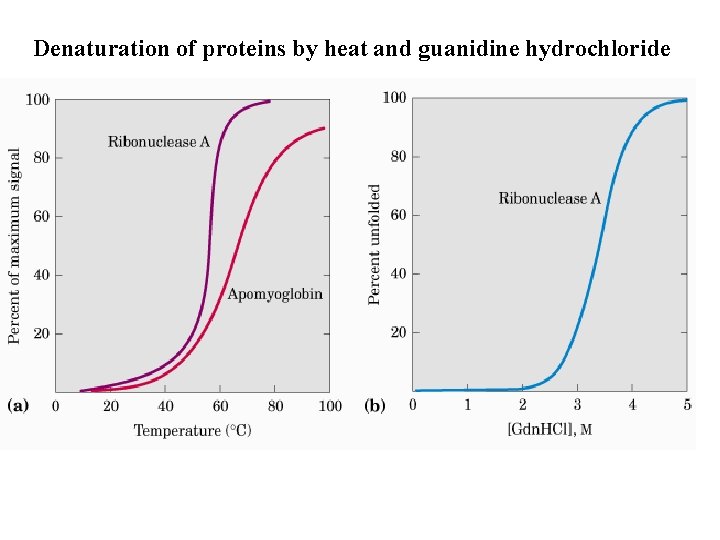
Ppt Denaturation Of Proteins By Heat And Guanidine Hydrochloride Powerpoint Presentation Id



Destruction And Building Of The Secondary And Tertiary




Dynamic Surface Properties Of Lysozyme Solutions Impact Of Urea And Guanidine Hydrochloride Sciencedirect




Protein Folding Under Mechanical Forces A Physiological View Physiology
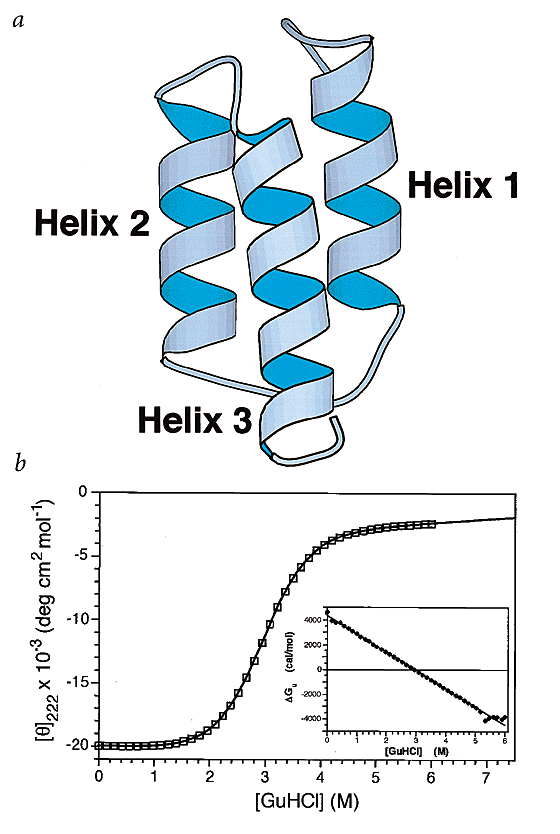



Preorganized Secondary Structure As An Important Determinant Of Fast Protein Folding Nature Structural Molecular Biology




A New Approach To Study The Physical Stability Of Monoclonal Antibody Formulations Dilution From A Denaturant Journal Of Pharmaceutical Sciences
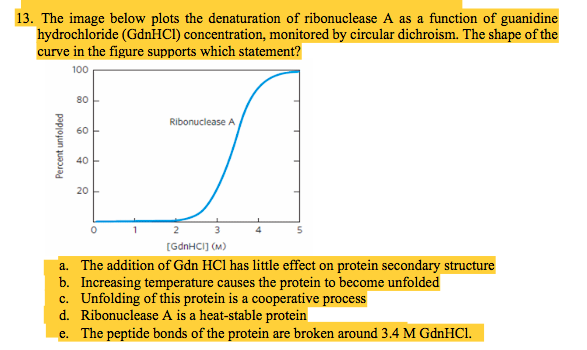



Solved Multible Choices 12 A Protein Will Chegg Com




Denaturation Of Protein Denaturation Denaturation Is A Process




Guanidine Is The Biochemical Compound It Is A Colourless Solid That Dissolves In Polar Solvents Stock Vector Vector And Low Budget Royalty Free Image Pic Esy Agefotostock




Guanidine Hcl
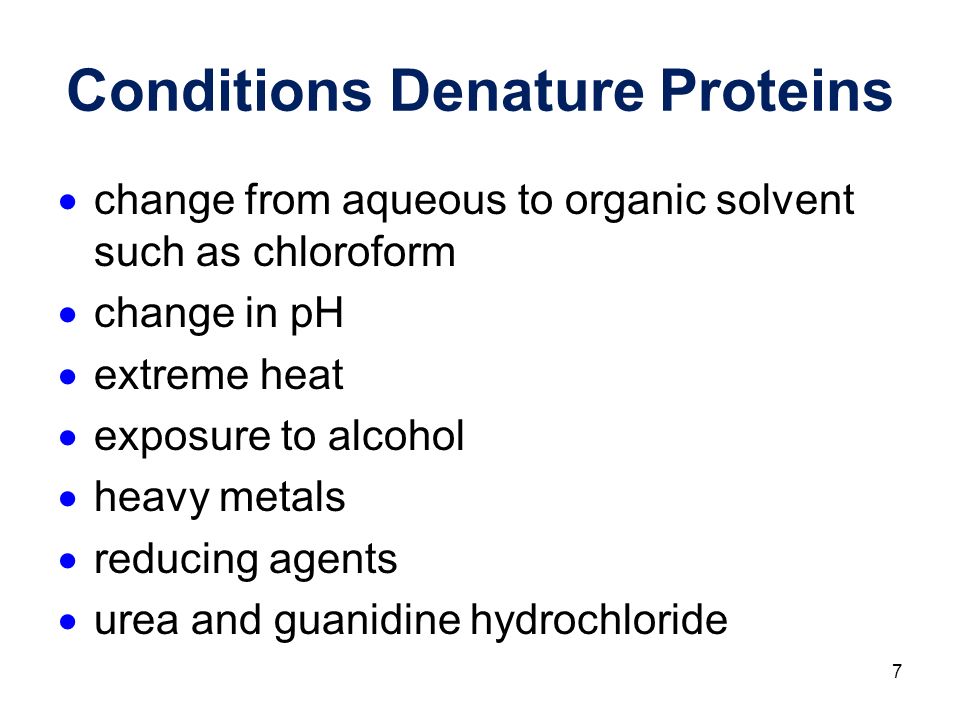




1 Outcome 2 Keeping Proteins In Their Native States Ppt Download




Preparation And Extraction Of Insoluble Inclusion Body Proteins From Escherichia Coli Palmer 1995 Current Protocols In Protein Science Wiley Online Library



Add Your Page Title



Add Your Page Title




Guanidine Compound Formula Hncnh22 Colourless Solid Stock Vector Royalty Free
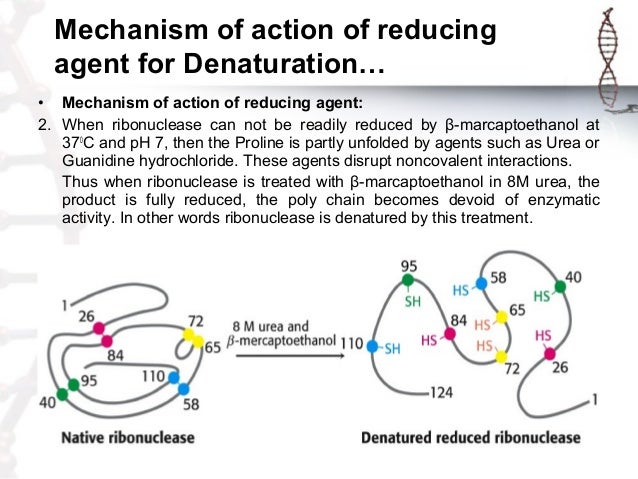



Protein




The Stability Of A Three State Unfolding Protein Intechopen




Investigation Of The Mechanism Of Protein Denaturation By Guanidine Hydrochloride Youtube



2
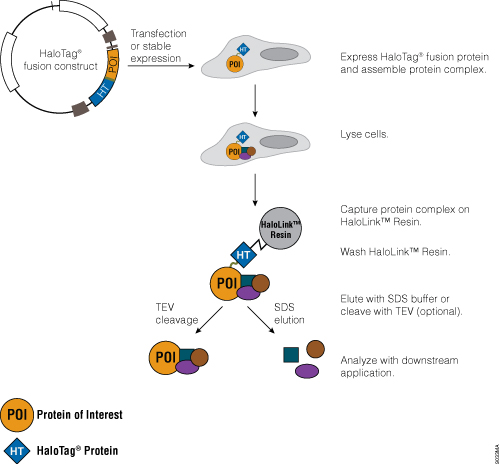



Protein Purification Guide An Introduction To Protein Purification Methods




Kinetic Analysis Of Guanidine Hydrochloride Inactivation Of B Galactosidase In The Presence Of Galactose




Preparation And Extraction Of Insoluble Inclusion Body Proteins From Escherichia Coli Abstract Europe Pmc



1




Enzymatic Degradation Of Rna Causes Widespread Protein Aggregation In Cell And Tissue Lysates Embo Reports



Minerva Access Unimelb Edu Au




Urea But Not Guanidinium Destabilizes Proteins By Forming Hydrogen Bonds To The Peptide Group Pnas




Pdf Urea And Guanidine Hydrochloride Denaturation Of Ribonuclease Lysozyme Zhymotrypsin And Lactoglobulin Semantic Scholar




Protein Denaturation And Misfolding



Plos One Differences In The Pathways Of Proteins Unfolding Induced By Urea And Guanidine Hydrochloride Molten Globule State And Aggregates




Guanidine Hydrochloride 50 01 1 Tci America




Comparison Of The Guanidine Hydrochloride Denaturation Curves Of Four Download Scientific Diagram




Contrasting Effects Of Guanidinium Chloride And Urea On The Activity And Unfolding Of Lysozyme Abstract Europe Pmc



Plos One Inactivation And Unfolding Of Protein Tyrosine Phosphatase From Thermus Thermophilus Hb27 During Urea And Guanidine Hydrochloride Denaturation
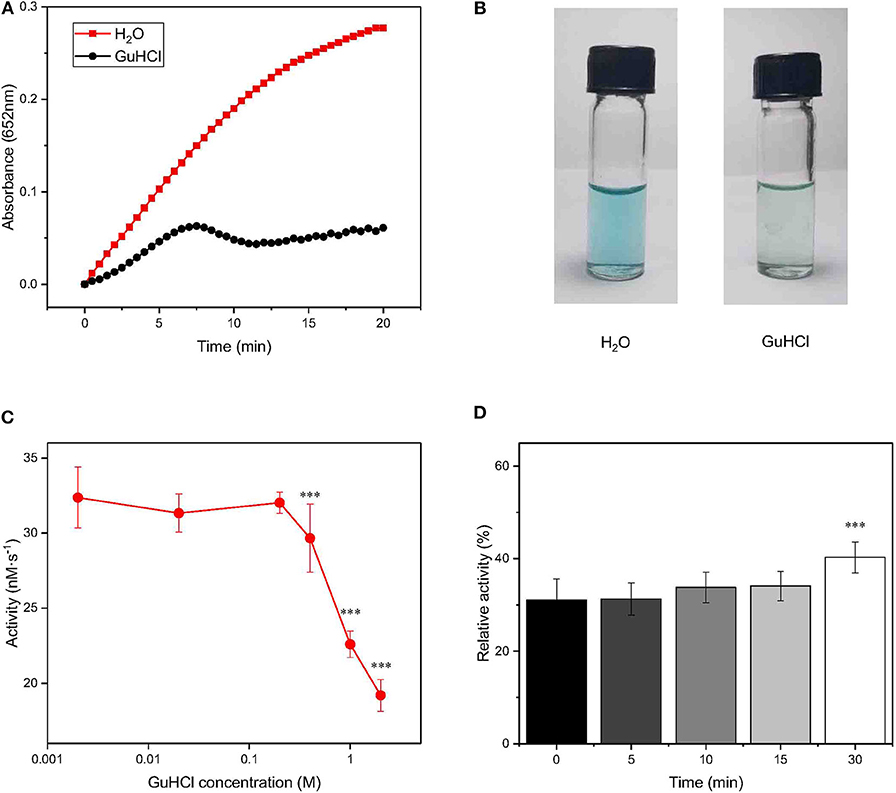



Frontiers Reversible Inhibition Of Iron Oxide Nanozyme By Guanidine Chloride Chemistry



Guanidine Hydrochloride 100 G Reagents For Protein Isolation Protein Isolation Biochemistry Life Science Carl Roth International




Solved Which Of The Following Are True About Guanidinium Chegg Com



What Are The Reagents Used In Denaturing Proteins Quora
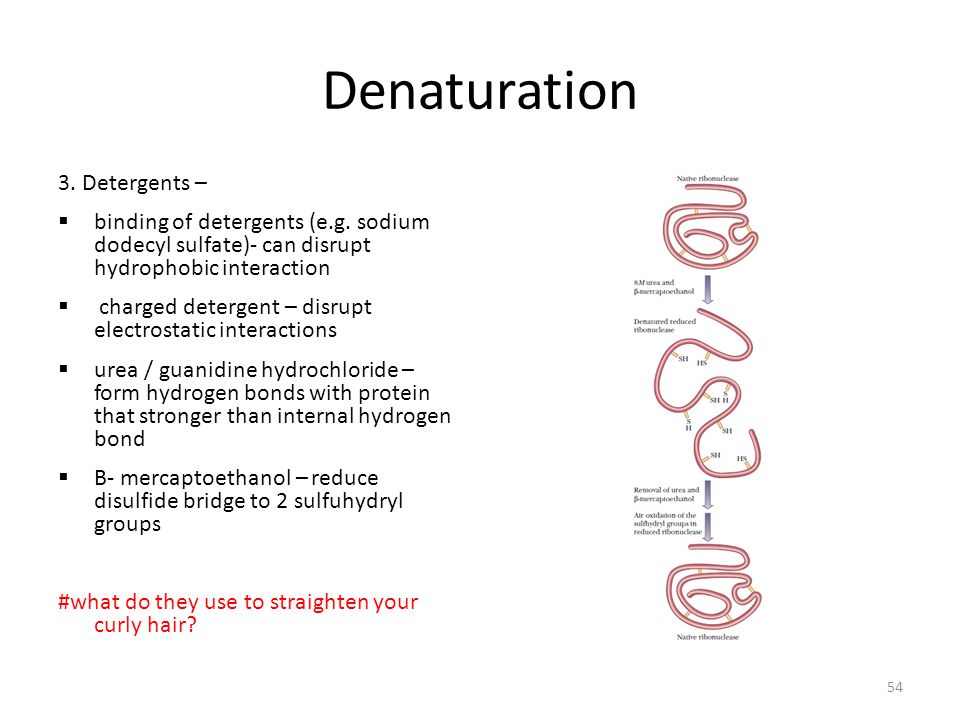




Topic 2 Biomolecule 1 Dna Protein Ppt Video Online Download




Gdn Hcl Millipore Bioz Ratings For Life Science Research



E Laboratory Determination




Comparative Refolding Of Guanidinium Hydrochloride Denatured Bovine Serum Albumin Assisted By Cationic And Anionic Surfactants Via Artificial Chaperone Protocol Biophysical Insight Sciencedirect
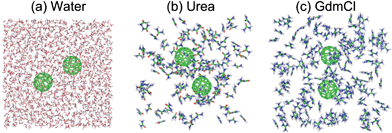



Dissociation Of Hydrophobic And Charged Nano Particles In Aqueous Guanidinium Chloride And Urea Solutions A Molecular Dynamics Study Nanoscale Rsc Publishing




Dodine As A Transparent Protein Denaturant For Circular Dichroism And Infrared Studies Guin 16 Protein Science Wiley Online Library



Guanidine Hydrochloride
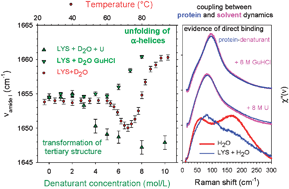



Influence Of Urea And Guanidine Hydrochloride On Lysozyme Stability And Thermal Denaturation A Correlation Between Activity Protein Dynamics And Conformational Changes Physical Chemistry Chemical Physics Rsc Publishing



Denaturation Biochemistry Wikipedia




Denaturation Of Glycinin By Urea And Guanidine Hydrochloride An Equilibrium And Kinetic Study Pdf Document




Enhanced Protein Mass Spectrometry Analysis With Trypsin Lys C Mix Promega Connections




Protein Stiffening And Entropic Stabilization In The Subdenaturing Limit Of Guanidine Hydrochloride Biophysical Journal
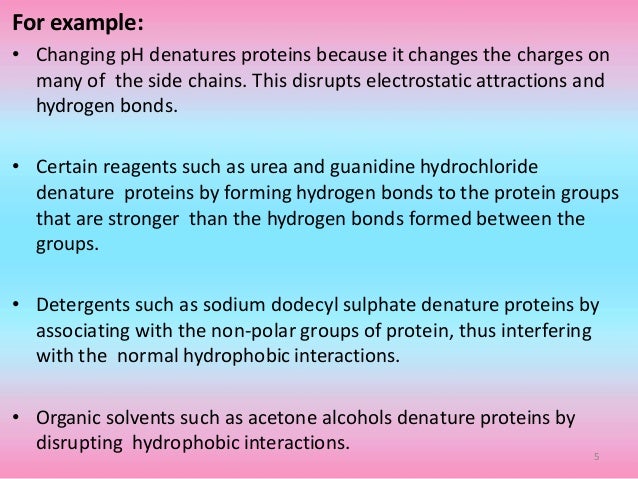




Protein Denaturation



D Protein Folding Stability In Vivo And In Vitro



2




Guanidine Hydrochloride Guhcl Induced Denaturation Of Wild Type And Download Scientific Diagram
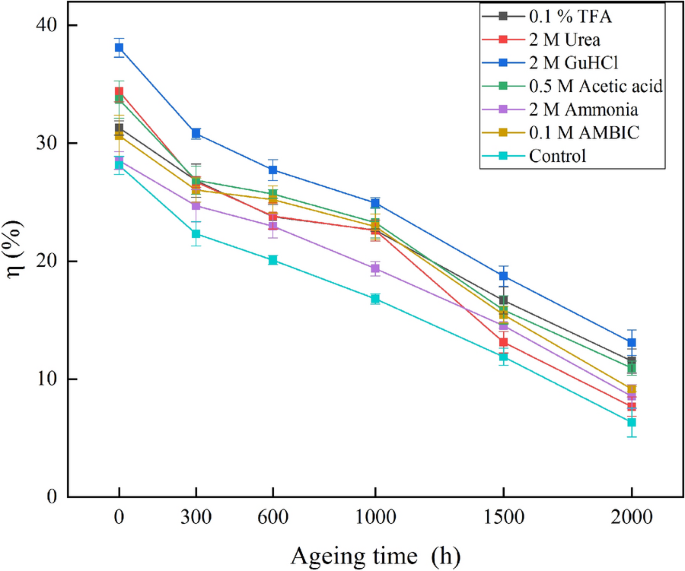



A Comparative Study On The Extraction Effects Of Common Agents On Collagen Based Binders In Mural Paintings Heritage Science Full Text




Denaturation Of The Wild Type And Mutant Bgltm Proteins With Guanidine Download Scientific Diagram




Guanidine Hydrochloride Cas 50 01 1 Ag Scientific Inc




Protein Aggregation Capture On Microparticles Enables Multipurpose Proteomics Sample Preparation Molecular Cellular Proteomics
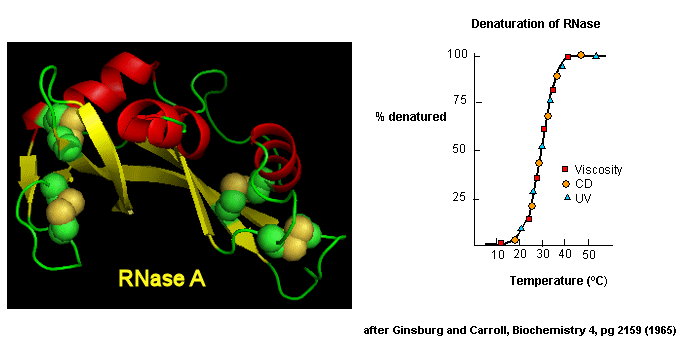



D2 Protein Folding In Vitro Biology Libretexts




Reversible Thermal Denaturation Of Human Fgf 1 Induced By Low Concentrations Of Guanidine Hydrochloride Biophysical Journal




Urea And Guanidinium Chloride Denature Protein L In Different Ways In Molecular Dynamics Simulations Biophysical Journal




Investigation Of Guanidine Hydrochloride Induced Chlorophyll Protein 43 And 47 Denaturation In The Terahertz Frequency Range Journal Of Applied Physics Vol 102 No 7




Guanidine Hydrochloride Denaturation Of Sc S100a4 Proteins A Download Scientific Diagram




Guanidine Hcl




Spectroscopic Studies On Unfolding Processes Of Apo Neuroglobin Induced By Guanidine Hydrochloride And Urea




Kinetic Evidence For A Two Stage Mechanism Of Protein Denaturation By Guanidinium Chloride Pnas



1
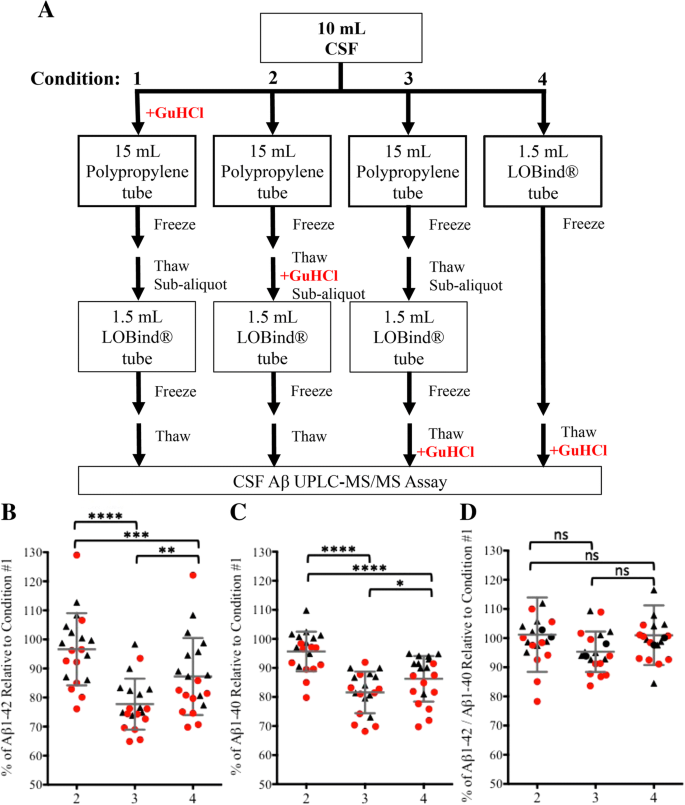



Preanalytical Approaches To Improve Recovery Of Amyloid B Peptides From Csf As Measured By Immunological Or Mass Spectrometry Based Assays Alzheimer S Research Therapy Full Text




Guanidine Is The Biochemical Compound It Is A Colourless Solid That Dissolves In Polar Solvents Stock Vector Vector And Low Budget Royalty Free Image Pic Esy Agefotostock
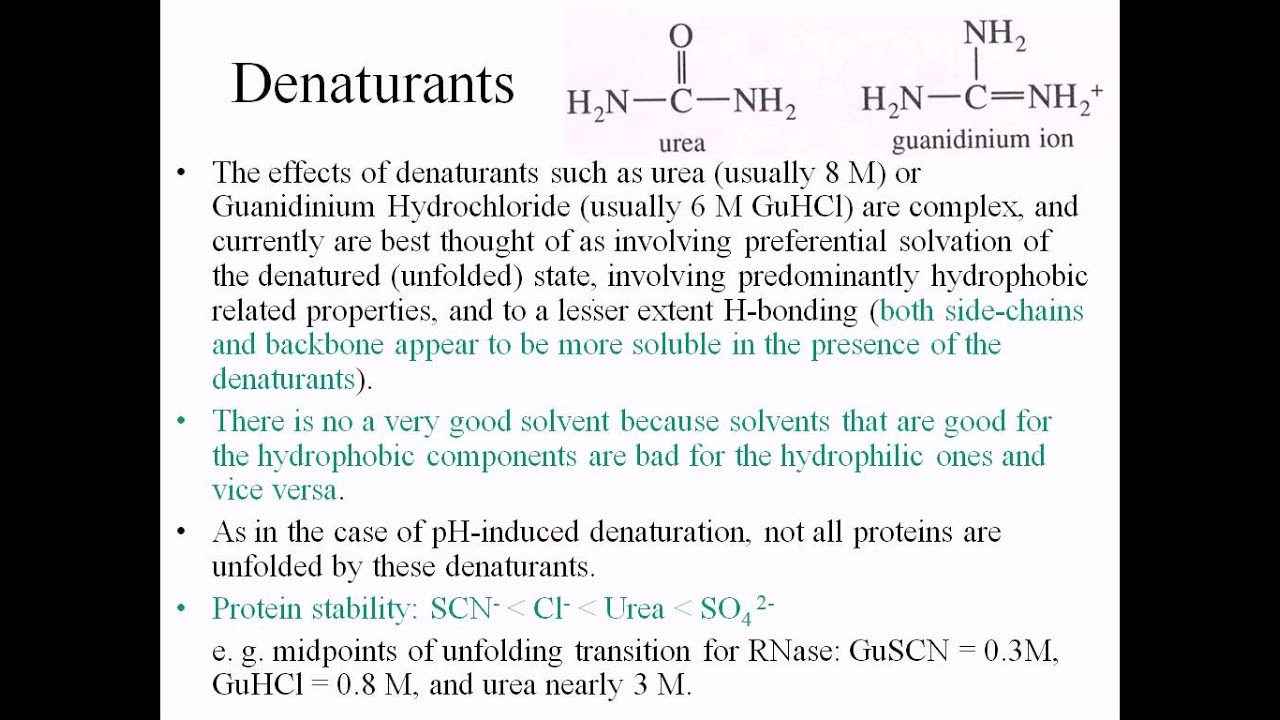




Protein Denaturing Agents Youtube




Biomolecules Free Full Text Interactions Under Crowding Milieu Chemical Induced Denaturation Of Myoglobin Is Determined By The Extent Of Heme Dissociation On Interaction With Crowders Html
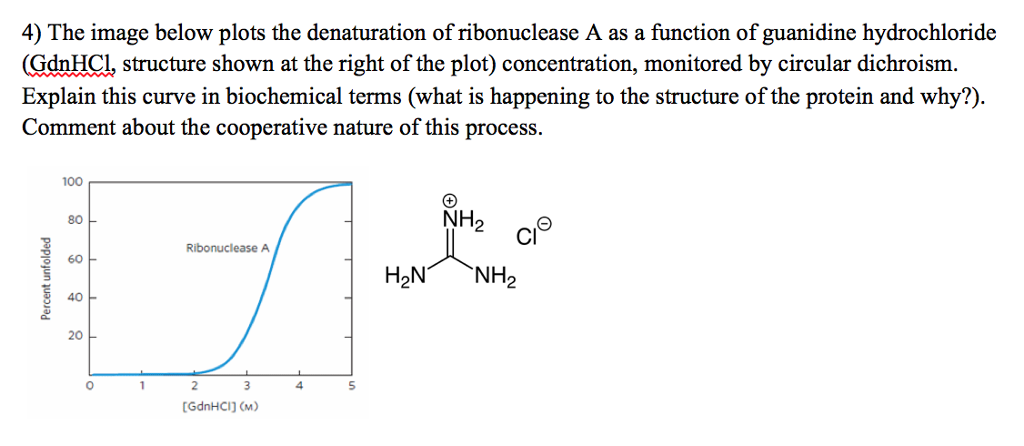



Solved The Image Below Plots The Denaturation Of Chegg Com




Guanidine Hydrochloride Guanidine Hcl Cas50 01 1 For Biology Research Buy Guanidine Hcl 50 01 1 Guanidine Thiocyanate Product On Alibaba Com




Action Of Guanidine Hydrochloride On Proteins




Detergents Triton X 100 Tween And More




The Ph Dependence Of Saccharides Influence On Thermal Denaturation Of Two Model Proteins Supports An Excluded Volume Model For Stabilization Generalized To Allow For Intramolecular Electrostatic Interactions Journal Of Biological Chemistry



0 件のコメント:
コメントを投稿